This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison. http://genetics564.weebly.com/
Protein Domains
What are protein domains and motifs?
Protein domains are areas of proteins that are highly conserved across species. One domain can also be present in a wide array of proteins. Protein domains are responsible for a particular function or interaction in the protein.
PTPN2 Domains
Pfam
Pfam is a database of conserved protein domain families that is used to annotate and classify proteins. The Y_phosphatase domain makes up the protein tyrosine phosphatase removes phosphate groups from tyrosine residues and can be involved in a wide array of cell interactions [1].
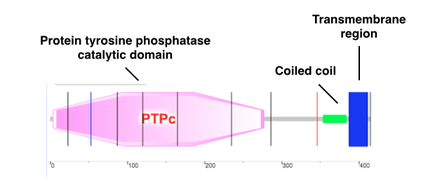
SMART
SMART (a Simple Modular Architecture Research Tool) is an online tool that identifies genetically mobile domains in specific proteins. The domains found in PTPN2 via SMART were the protein tyrosine phosphatase catalytic domain (PTPc), a coiled coil, and a transmembrane region. Protein tyrosine phosphatase is a postranslational modification that catalyses the removal of a phosphate group attached to the tyrosine residue. PTPc is involved in many cellular functions like cellular localization, affect protein stability, and regulate enzyme activity. The coiled coil is a structural motif. The transmembrane region is located at the end of the sequence and allows binding.
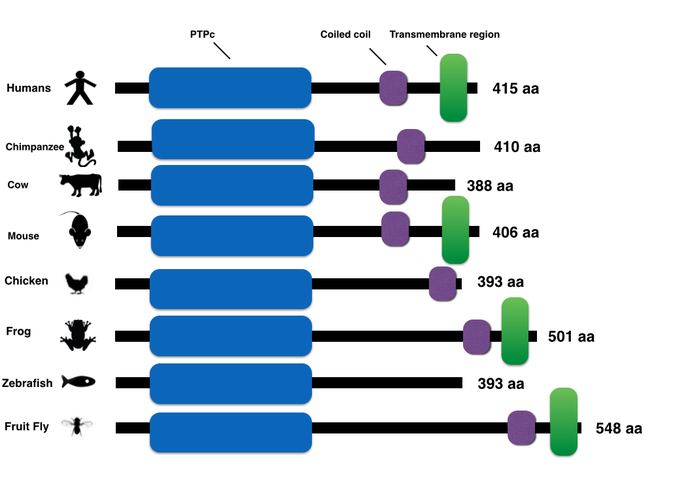
PTPN2 domain comparison with other model organisms
Analysis
The domains within PTPN2 are a crucial part of determining the protein's function. The most important domain in PTPN2, the catalytic protein tyrosine phosphatase domain (PTPc), is important because the dysfunction in the Crohn's Disease PTPN2 mutant is located within this domain. The PTPc domain is also a vital piece to PTPN2's function in dephosphorylating proteins, which is what I will be further investigating in my specific aims.
Conservation of protein domains among different organisms is also important to note when utilizing protein homologs. For example, the zebrafish is lacking the transmembrane domain in PTPN2. This missing region can have potential implications for varying protein localization between zebrafish and humans and can alter how conclusive your results can be.
Conservation of protein domains among different organisms is also important to note when utilizing protein homologs. For example, the zebrafish is lacking the transmembrane domain in PTPN2. This missing region can have potential implications for varying protein localization between zebrafish and humans and can alter how conclusive your results can be.
References
- http://pid.nci.nih.gov/2011/110913/full/pid.2011.3.shtml
- http://smart.embl-heidelberg.de/help/smart_about.shtml